PDF Raycheva Tz., Stoyanov K., & Denev I. 2011.
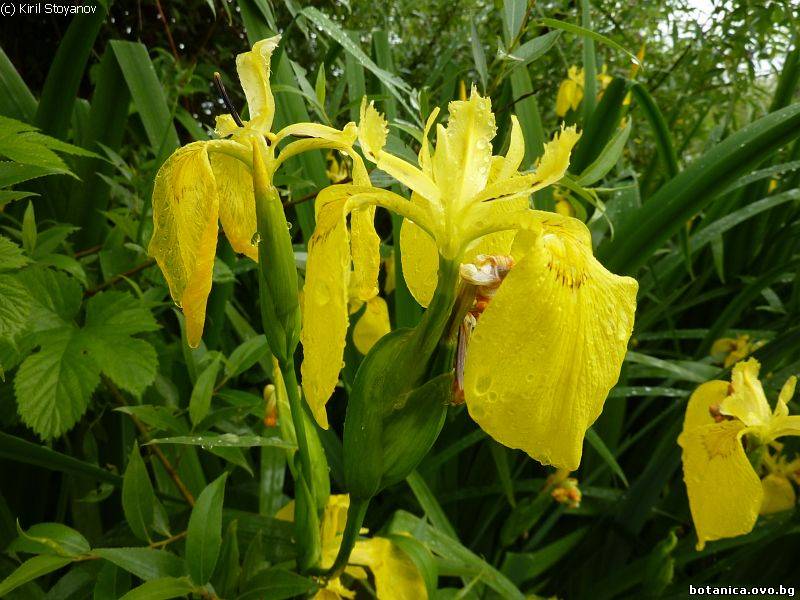
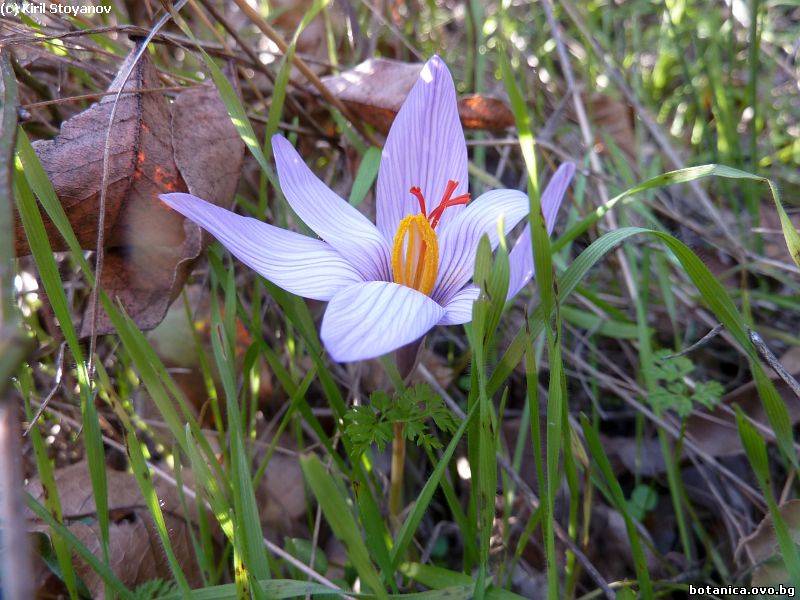
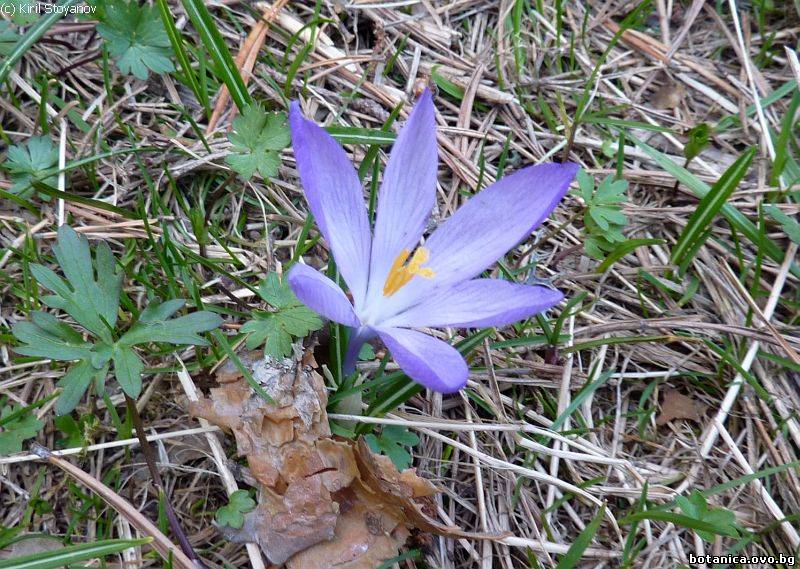
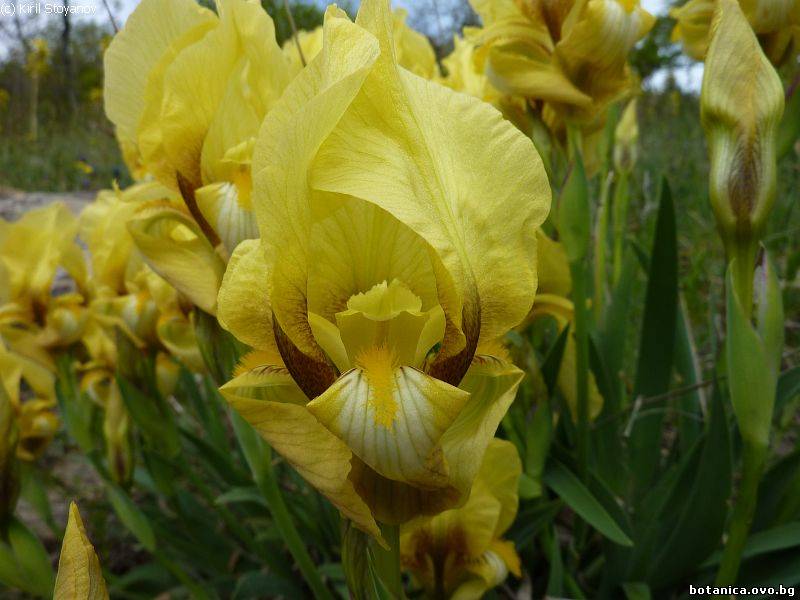
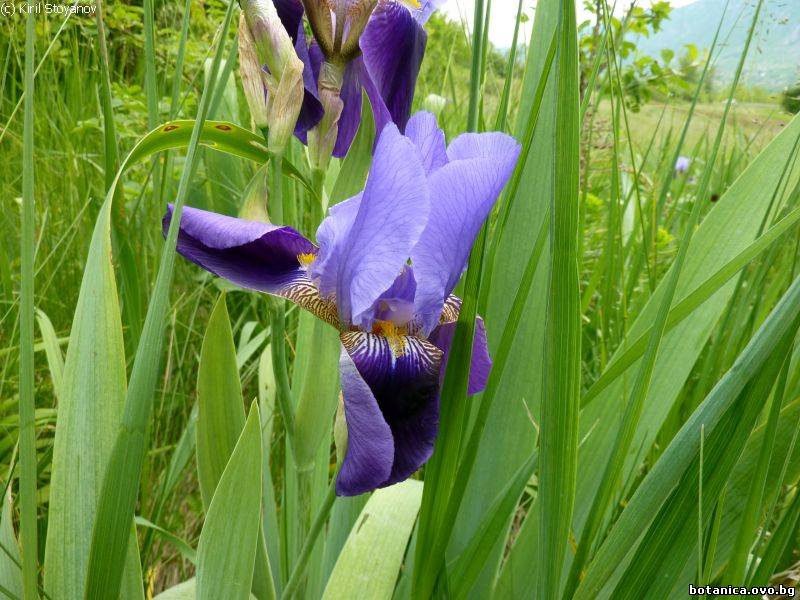
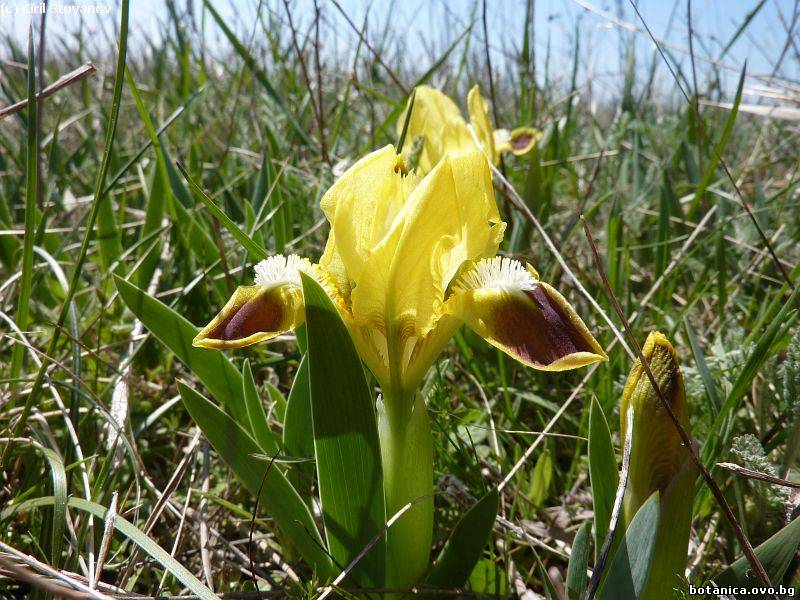
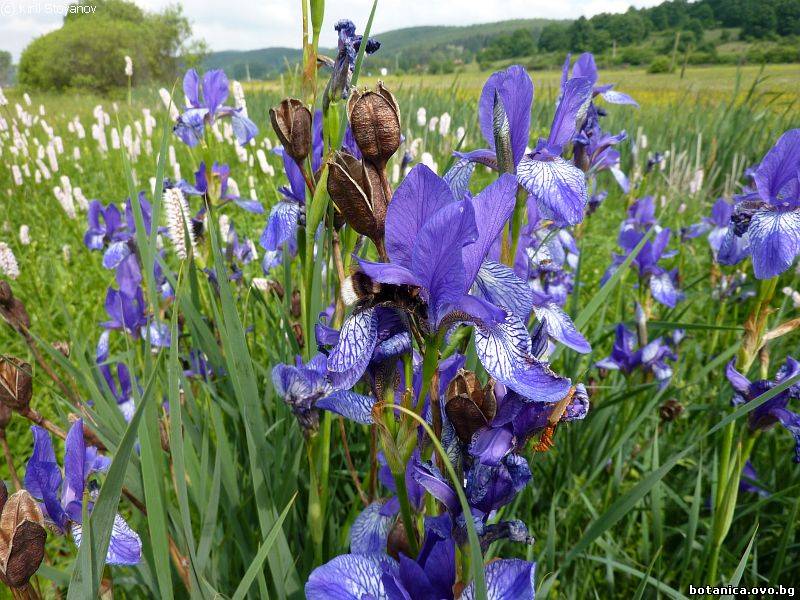
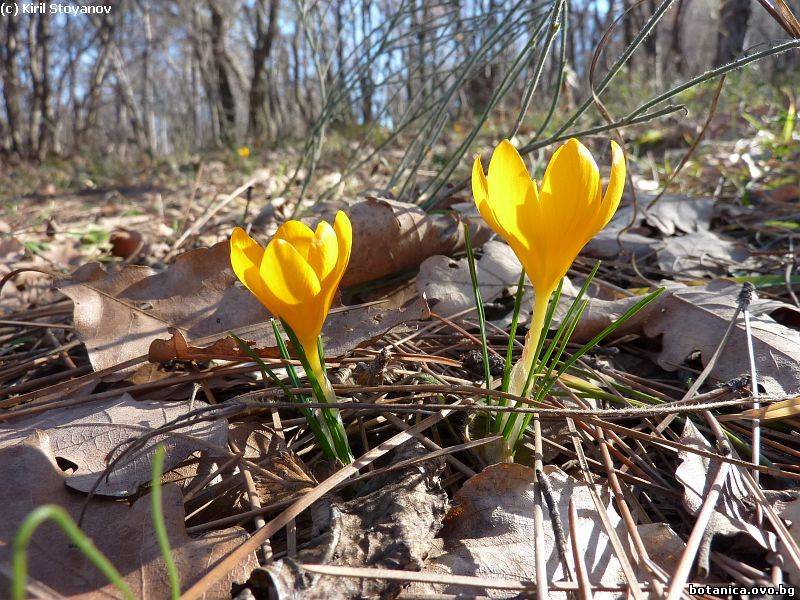
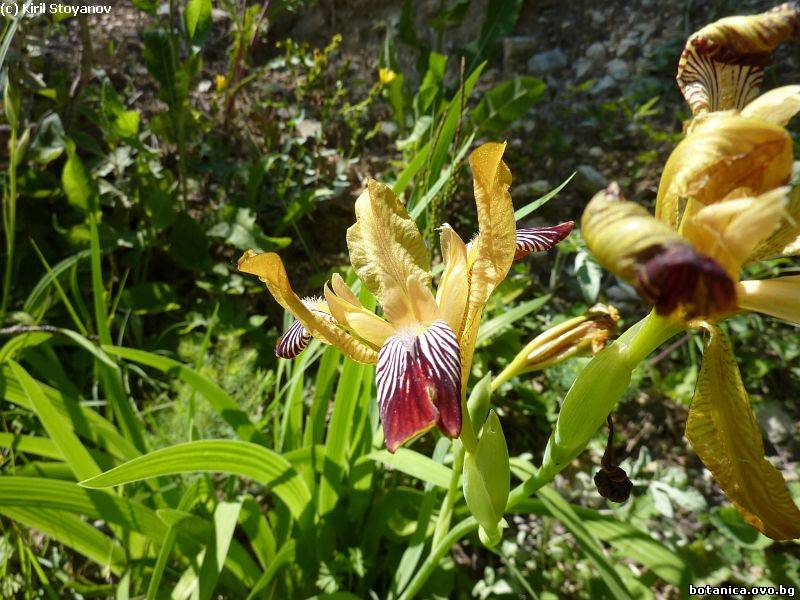
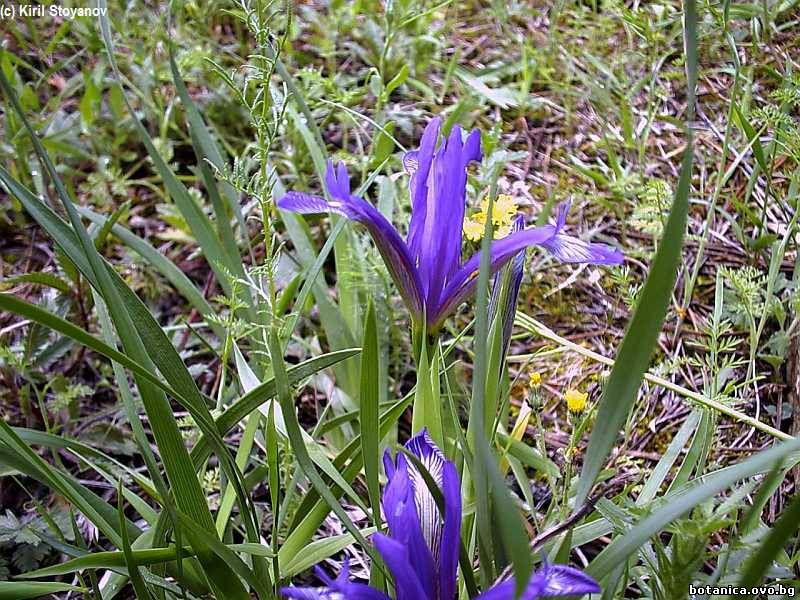
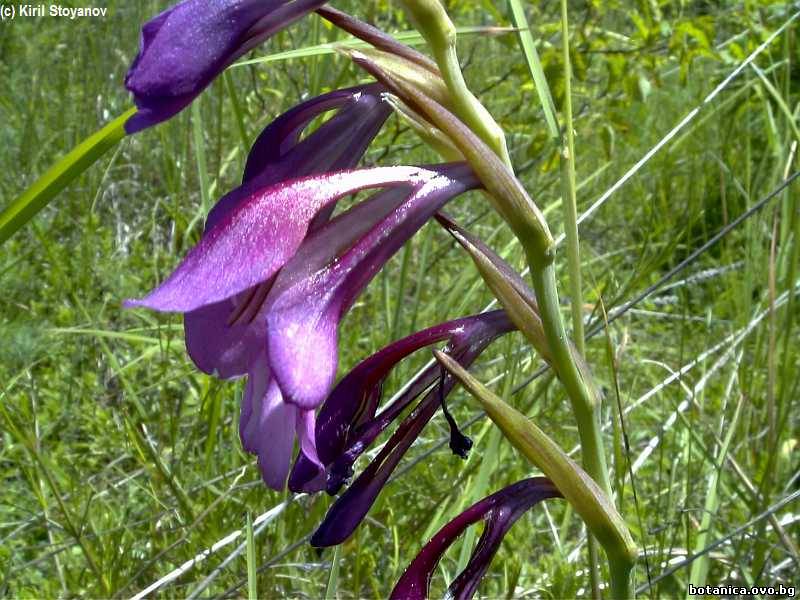
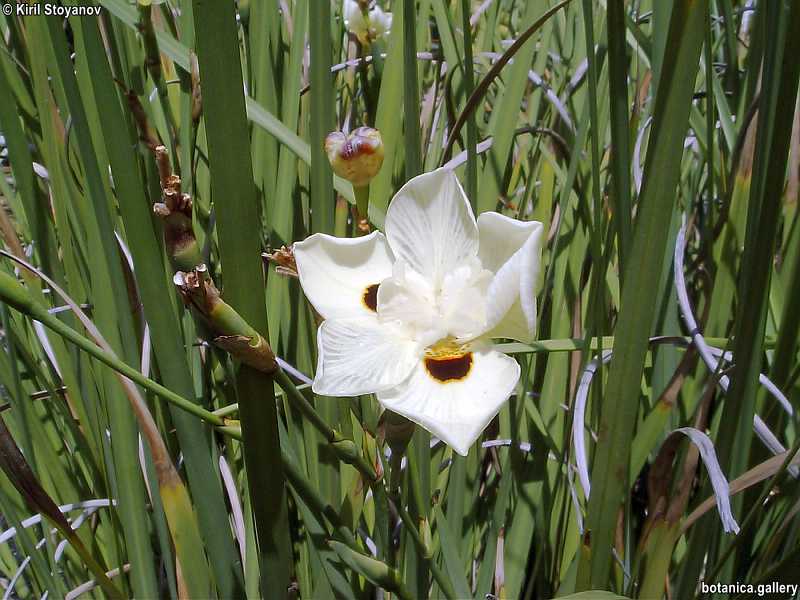
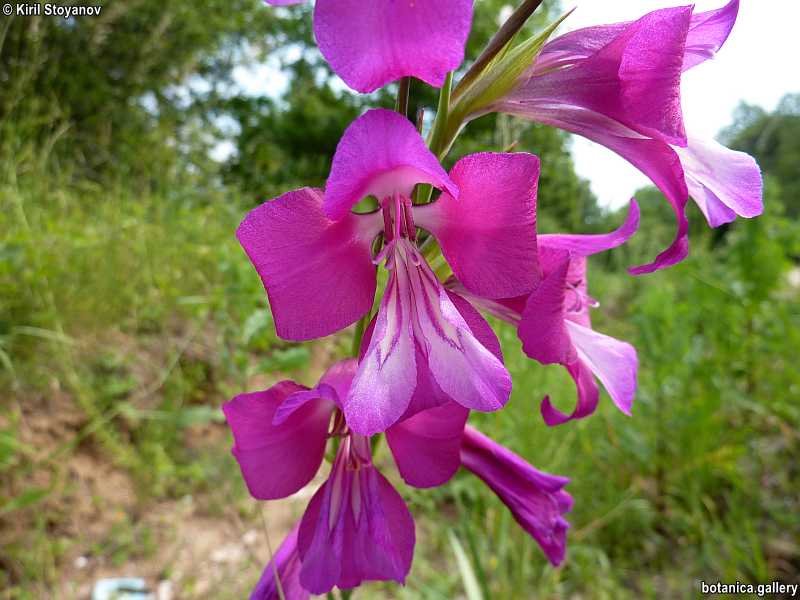
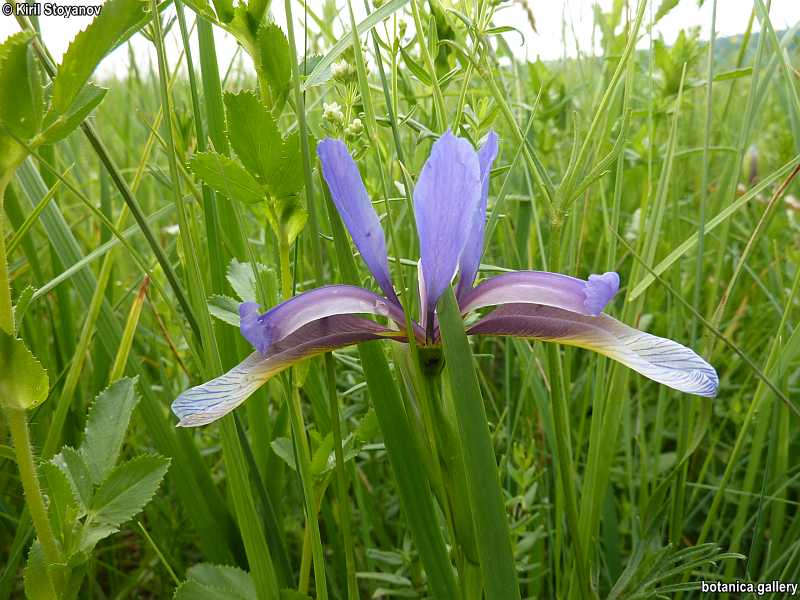
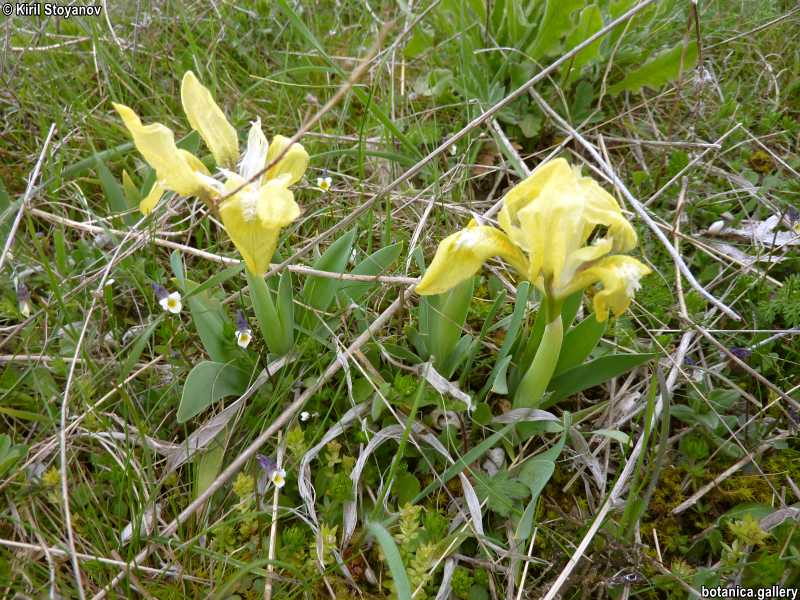
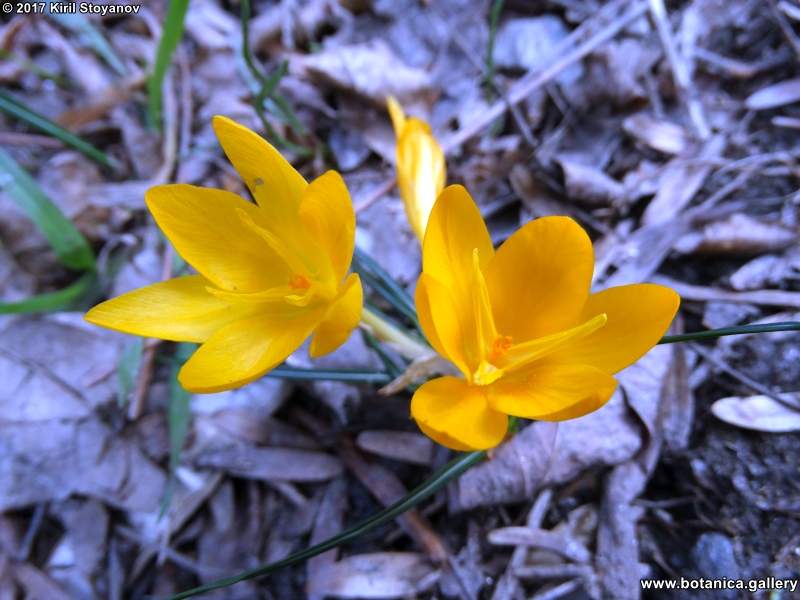
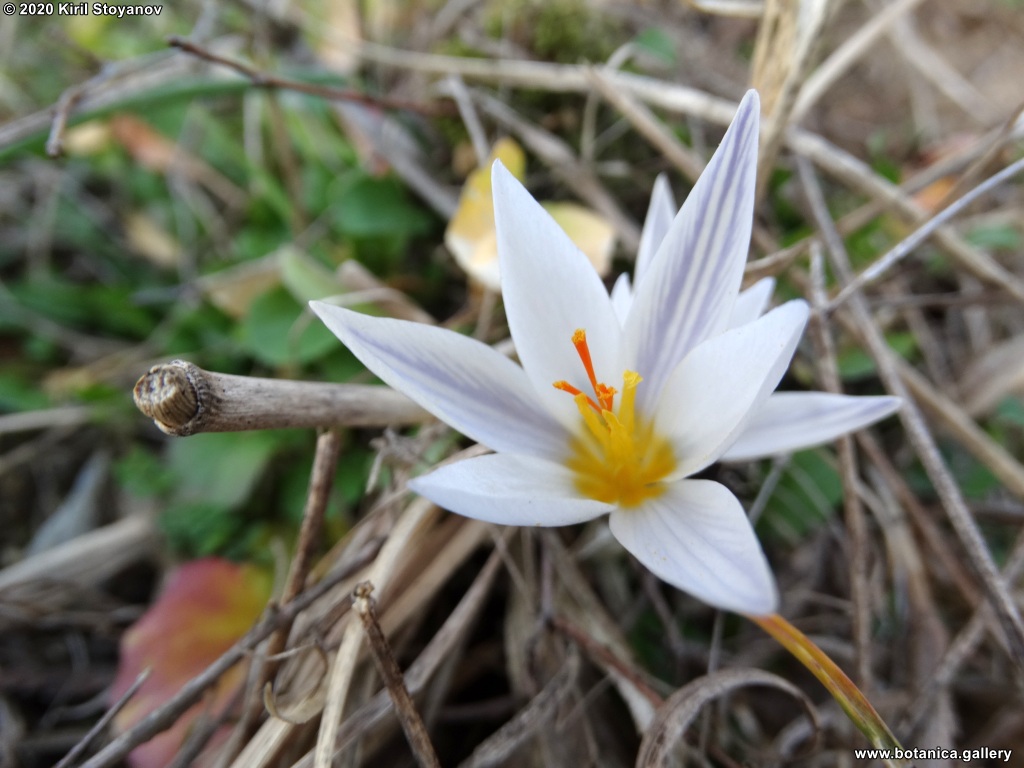
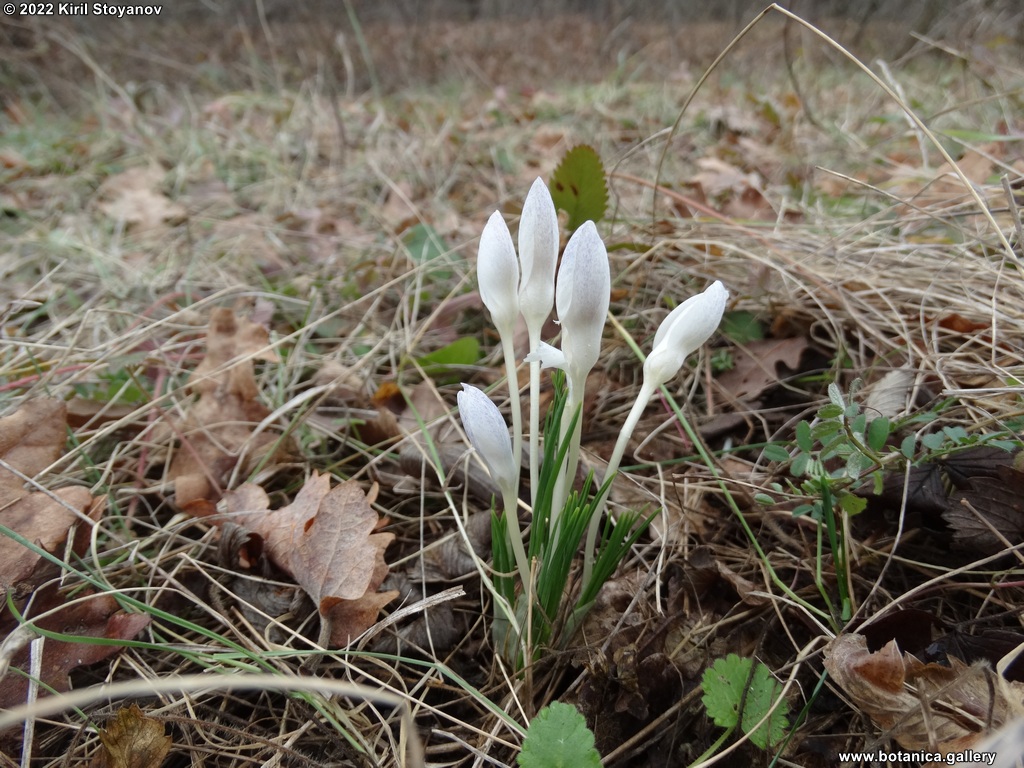
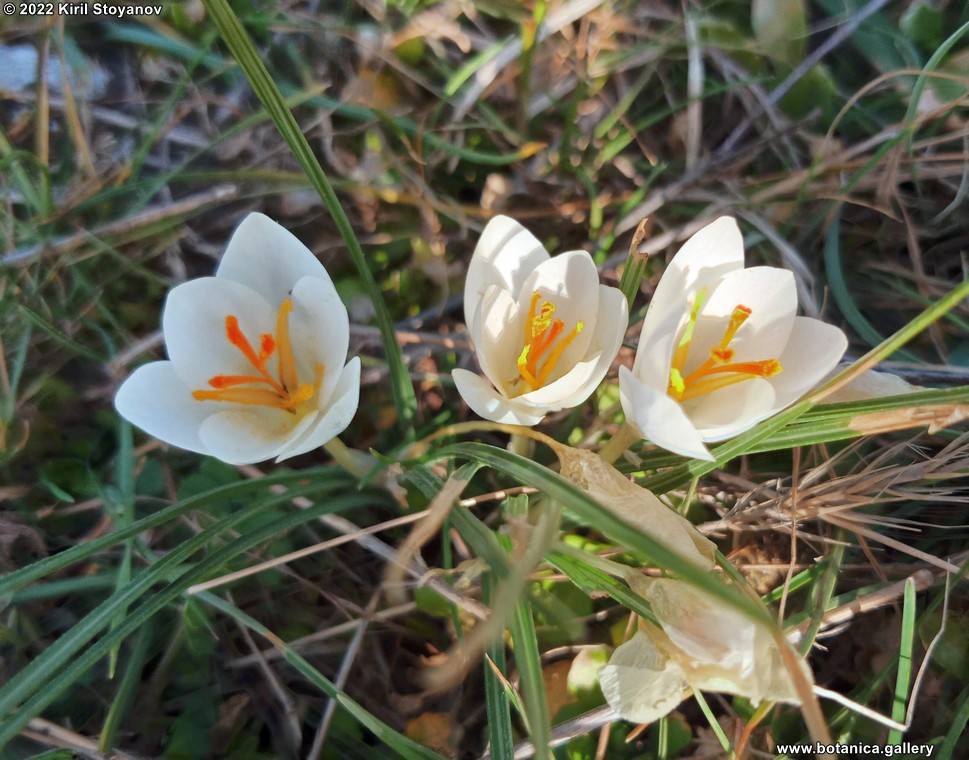
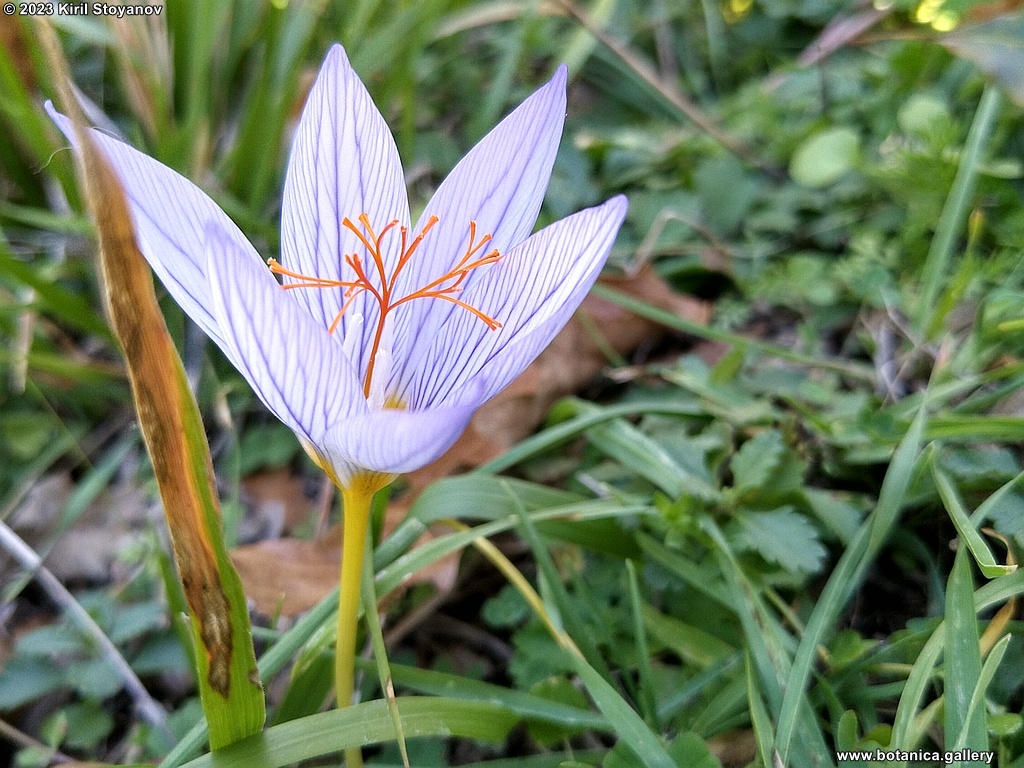
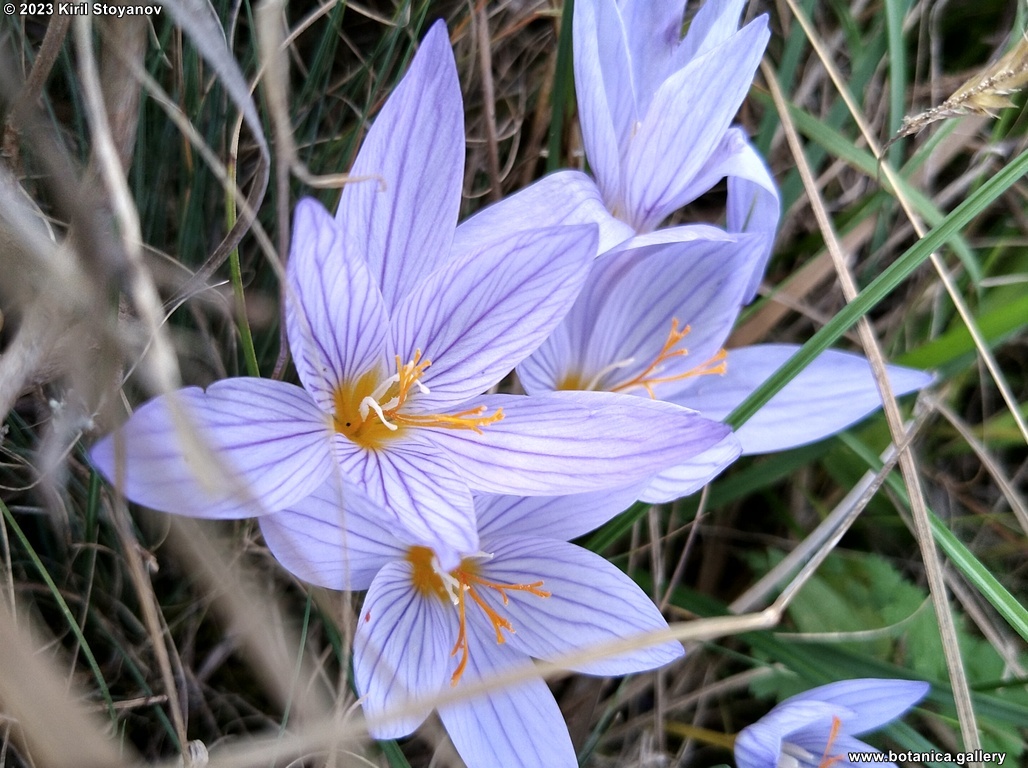
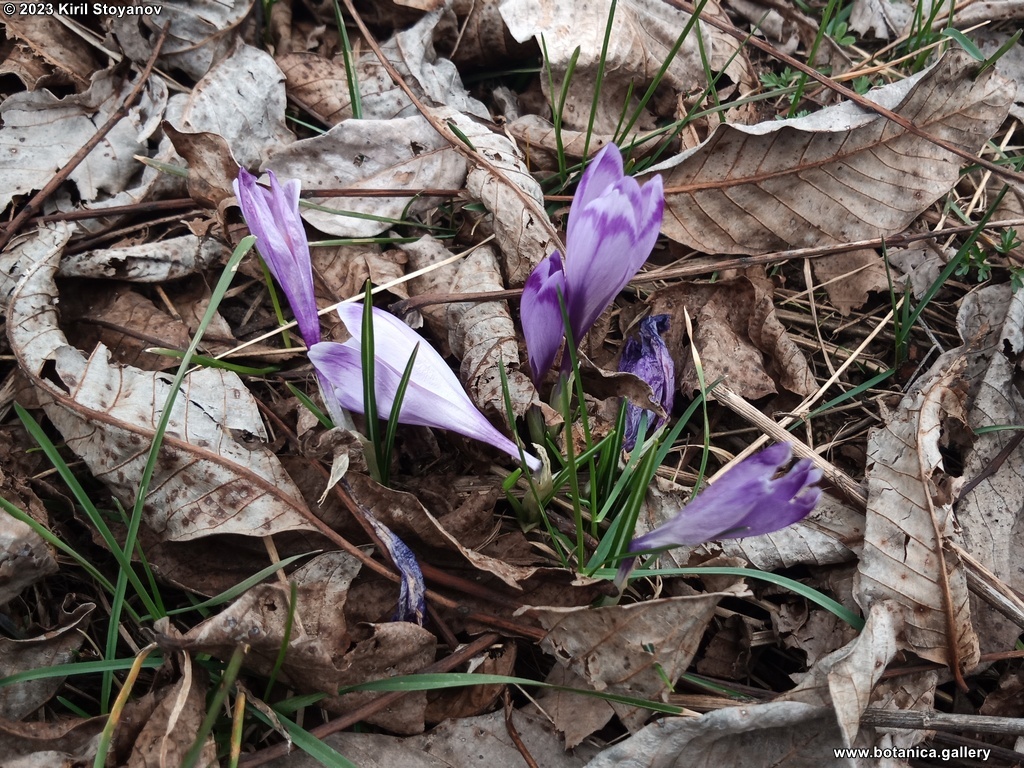
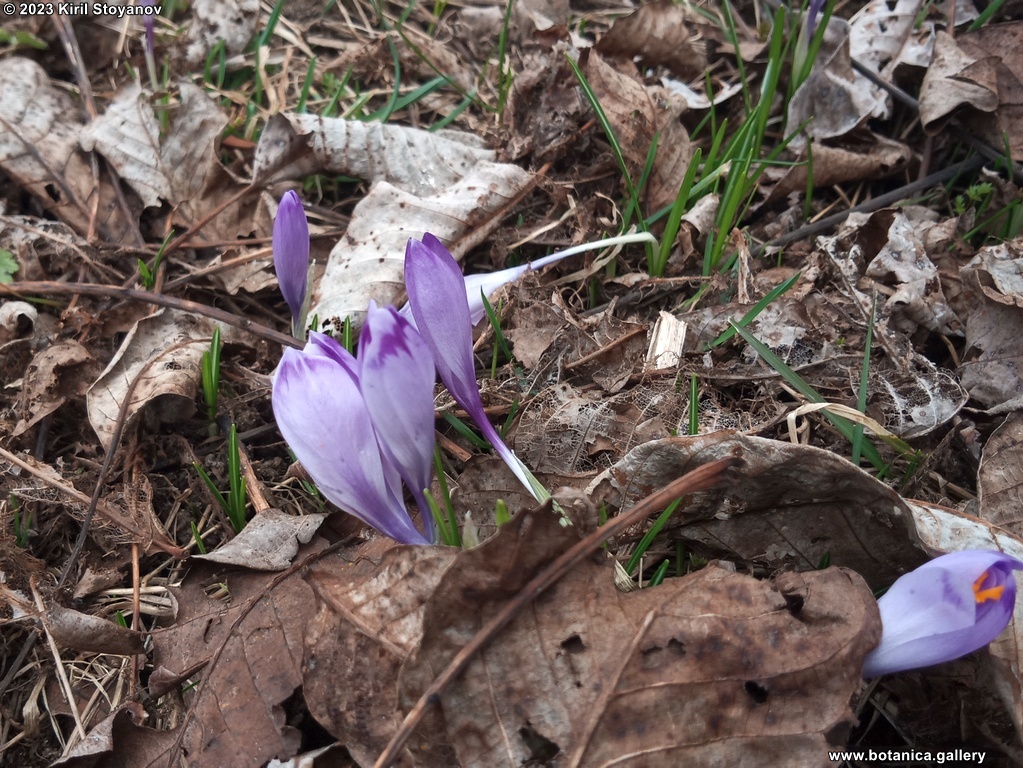
Genetic diversity and molecular taxonomy study of three genera from Iridaceae family in the Bulgarian flora based on ISSR markers
Biotechnology & Biotechnological Equipment 25(3): 2484-2488.
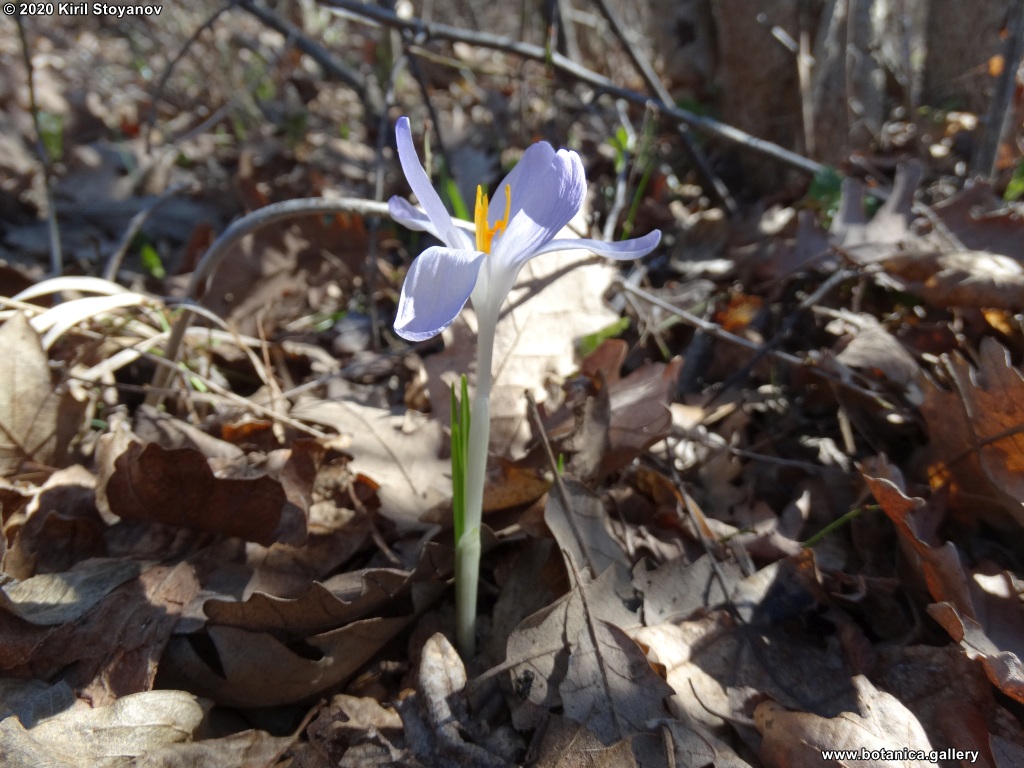
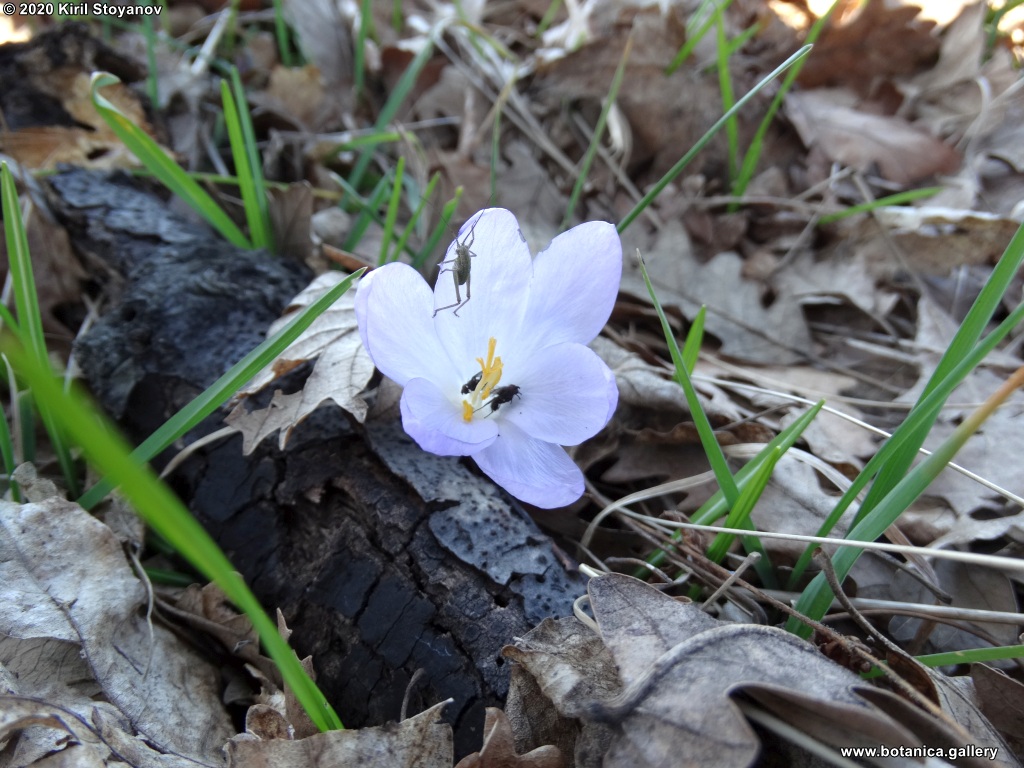
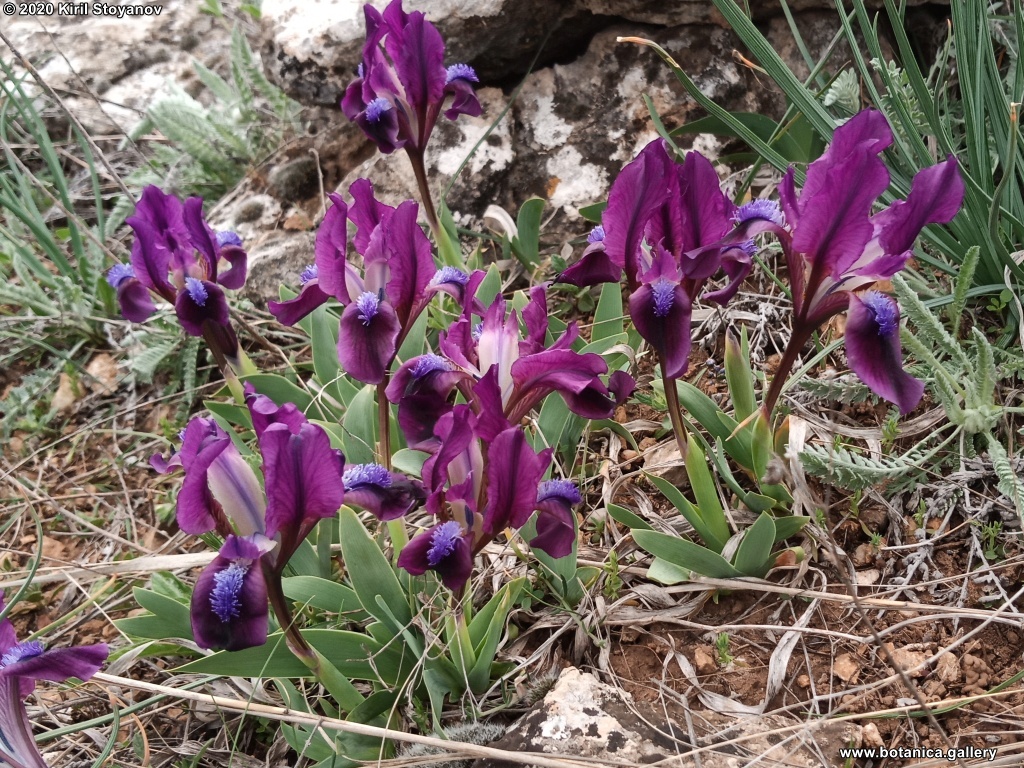
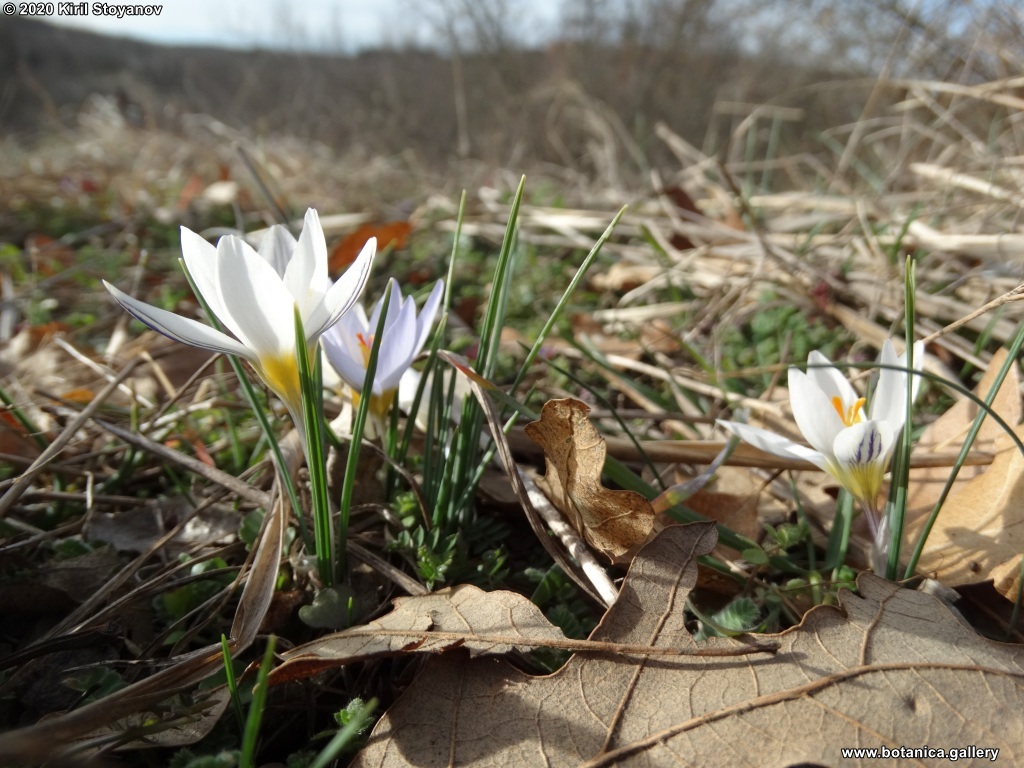
Abstract: Iridaceae is a family of perennial plants with almost worldwide distribution. The taxonomy of the family is based mainly on the morphology, anatomy, embryology and chromosome numbers. The systematics and phylogeny within the family is still subject of debates. This is mainly because most of the classification schemes and determination keys are based on morphological descriptions. These features are often unreliable for exact determination of the species with convergent morphology but inhabiting different ecological localities. In Bulgaria the family is represented by four genera. The genetic diversity and the relations between seven Bulgarian species from the Iridaceae family were examined by ISSR markers. The ISSR-PCR reactions were carried out with seven different ISSR primers and genomic DNA isolated from 13 samples. The distribution of polymorphic PCR products was analyzed by PAST software. The combined results of genetic variants were used to construct a consequent unrooted diagram.
The obtained results clearly defined the two subfamilies in Iridaceae family – Iridoideae and Crocoideae Burnett. The observed grouping of studied species did not coincide with the classification schemes based on morphology features, but was in agreement with the phylogenic studies. Our data confirmed the hypothesis for the polyphyletic origin of species from subgenus Limniris (Tausch) Spach – ser. Laevigatae (Diels) Lawrence and showed their genetic similarity to subgenus Iris ser. Iris. This work, to our best knowledge, is the first attempt to reassess by means of molecular markers the entire taxonomical scheme of the recent members of Iridaceae in Bulgaria.
Acknowledgements
the study was supported by the Scientific Research Center of Agricultural University – Plovdiv (grant 02-10) and the National Science fund of Bulgaria grants DТК 02/40 and BG051Po001-3.3.04/17., nAto grant clG 983884 and ЕRA 117.
Scopus, Web of Science, IF 1.632
[цитирано в: | cited in:]
- Ahmed Z., Dhatt, K.K., Nazir N. 2017. Genetic diversity analysis and DNA fingerprinting of elite stock of Gladiolus (Gladiolus hybridus Hort.) using ISSR markers. Applied Biological Research, 19(3) 290-298. Web of Science, Scopus
- Crespo M.B., Martinez-Azorin M. & Mavrodiev E. 2015. Can a rainbow consist of a single colour? A new comprehensive generic arrangement of the ‘Iris sensu latissimo’ clade (Iridaceae), congruent with morphology and molecular data. Phytotaxa, 232 (1), DOI: http://dx.doi.org/10.11646/phytotaxa.232.1.1 Web of Science, Scopus
- Colasante M, Fadda A., Rudall P.J. & Tarquini. 2021. The genus Iris as a critical taxon in establishing an integrated approach to Italian plant biodiversity. Fl. Medit. 31 (Special Issue): 213-239. https://doi.org/10.7320/FlMedit31SI.213 Scopus
- Datta SK, 2020. Breeding of Ornamentals: Gladiolus. LS – An International Journal of Life Sciences, Vol. 9, No. 2, pp. 115-133.Scopus
- De Castro O., De Guacchio E., Di Iorio E., Di Maio A., Di Martino L., Bartolucci F., Conti F. 2020. Barcoding helps threatened species: the case of Iris marsica (Iridaceae) from the protected areas of the Abruzzo (Central Italy). Plant Biosystems. DOI: 10.1080/11263504.2020.1762786 Web of Science, Scopus
- Dhiman M.R., Thakur N., Gupta Y.C., Sharma N. 2022 Gladiolus. In: Datta S.K., Gupta Y.C. (eds) Floriculture and Ornamental Plants. Handbooks of Crop Diversity: Conservation and Use of Plant Genetic Resources. Springer, Singapore. https://doi.org/10.1007/978-981-15-1554-5_5-1
- Emna E.C., Faouzi H. 2015. Analysis of diversity among wild gladiolus (Gladiolus sp.) accessions using morphological traits. International Journal of Agronomy and Agricultural Research (IJAAR), 6(1): 54-62.
- Hiremath V., Singh K.P., Jain N., Swaroop K., Jain P.K., Panwar S. & Sinha N. 2023. Cross Species/Genera Transferability of SSR Markers, Genetic Diversity and Population Structure Analysis in Gladiolus (Gladiolus × grandiflorus L.) Genotypes. PeerJ 11(1):e15820, http://dx.doi.org/10.7717/peerj.15820
- Kumar, M., Sirohi, U., Yadav, M.K. & Chaudhary V. 2023. In Vitro Culture Technology and Advanced Biotechnology Tools Toward Improvement in Gladiolus (Gladiolus species): Present Scenario and Future Prospects. Mol Biotechnol. https://doi.org/10.1007/s12033-023-00818-8 Scopus
- Nosrati H., Movafeghi A., Saffar S., Feizi M.H., Haghighi A. 2014. Systematic Applicability of ISSR Markers at Intra-Familial Level. Analele Universităţii din Oradea, Fascicula Biologie. 21(1): 14-18.
- Okba M.M., Abdel Baki P.M., Khaleel A.E., El-Sherei M.M., Salem M.A. 2021. Discrimination of common Iris species from Egypt based on their genetic and metabolic profiling. Phytochemical Analysis. doi:10.1002/pca.2945 Web of Science, Scopus
- Singh N., Pal A.K., Roy R.K., Tewari S.K., Tamta S., Rana T.S. 2016. Assessment of genetic variation and population structure in Indian Gladiolus cultivars inferred from molecular markers. The Nucleus. 1–10. doi:10.1007/s13237-016-0181-4 Web of Science, Scopus
- Singh N., Meena B., Pal A.K., Roy R.K., Tewari S.K., Tamta S., Rana T.S. 2016. Nucleotide diversity and phylogenetic relationships among Gladiolus cultivars and related taxa of family Iridaceae. Journal of Genetics. DOI 10.1007/s12041-017-0755-1 [http://www.ias.ac.in/public/Resources/General/jgen/16-574-030317.pdf]
- Singh N. 2017. Development of ISSR- and RAPD-derived SCAR markers for identification of Gladiolus germplasm. Journal of Horticultural Science and Biotechnology. https://doi.org/10.1080/14620316.2017.1309995 Web of Sciences, Scopus
- Singh N., Pal A.K., Roya R.K., Tamtab S. & Ranaa T.S. 2017. Development of cpSSR markers for analysis of genetic diversity in Gladiolus cultivars.. Plant Gene, Volume 10, June 2017, Pages 31–36. https://doi.org/10.1016/j.plgene.2017.05.003 Web of Science, Scopus
- Singh N., Pal A.K., Roy R.K., Tewari S.K. & Rana T.S. 2017. Characterization of Gladiolus Germplasm Using Morphological, Physiological, and Molecular Markers. Biochem Genet. Web of Science https://doi.org/10.1007/s10528-017-9835-4 Web of Science, Scopus
- Singh N., Meena B., Pal A.K., Roy R.K., Tewari S.K., Tamta S. & Rana T.S. 2018. Nucleotide diversity and phylogenetic relationships among Gladiolus cultivars and related taxa of family Iridaceae Journal of Genetics, 96(1): 135–145.
https://doi.org/10.1007/s12041-017-0755-1 Web of Science, Scopus - Singh N., Mahar K.S. Verma S., Meena B., Roy R.K., Tewari S.K., Goel A.K. & Rana T.S. 2018. Molecular analysis of genetic variability and relationship among Gladiolus cultivars. Indian Journal of Biotechnology, 17: 118-127. Web of Science, Scopus
- Zangeneh M. & Salehi H. 2019. Genetic Diversity as Revealed by Intersimple Sequence Repeat Polymorphism in Narcissus Accessions to Identify the Tolerant Genotypes for Deficit Irrigation. Journal of the American Society for Horticultural Science, 144(2): 92-106. https://doi.org/10.21273/JASHS04583-18 Web of Science, Scopus